What is ReproZip?
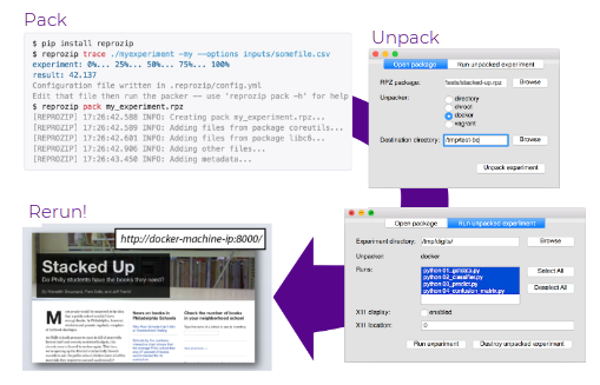
ReproZip is an open source tool that helps users make their work completely reproducible. ReproZip can be used to reproduce a plethora of applications, including data analysis tools, scripts and software written in any programming language, graphical tools, interactive tools, client-server applications (including databases), Jupyter notebooks, and more!
ReproZip works by tracing the systems calls used by the experiment to automatically identify which files should be included. Users can review and edit this list and the metadata before creating the final package file. Packages can be reproduced in different ways, including chroot environments, Vagrant-built virtual machines, and Docker containers; more can be added through plugins. We are in the process of merging in a new unpacker for Singularity right now!
You can also view a few playlists we've compiled that demo how to pack and unpack different applications with ReproZip:
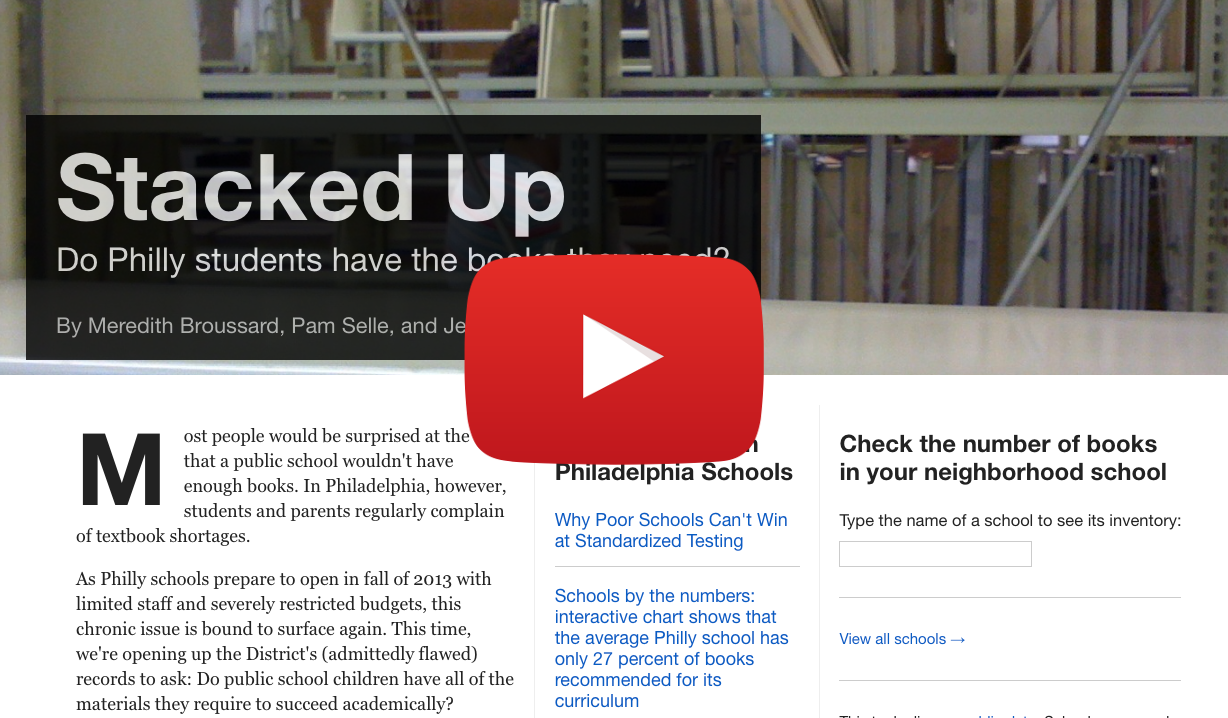
Stacked Up
Dynamic website called Stacked Up, which was created as a data journalism project to assess whether kids in Philly schools have the books they need.
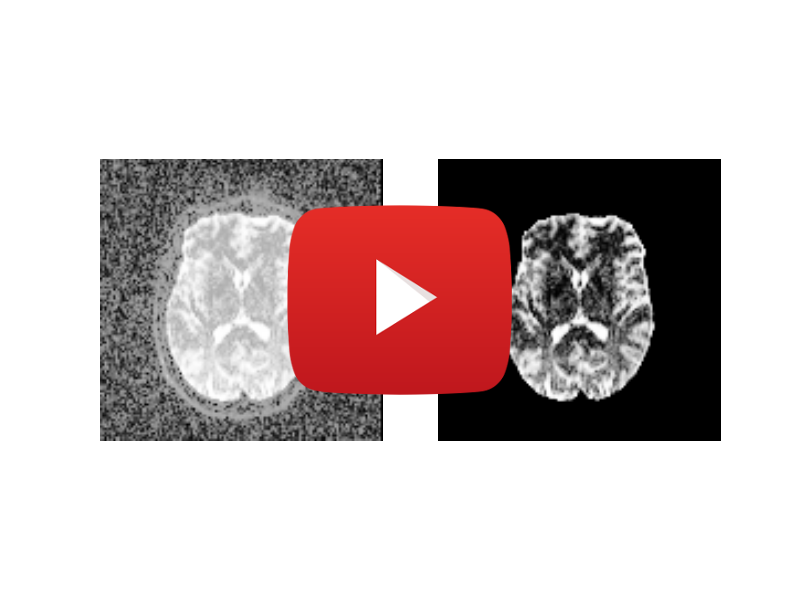
Brain Segmentation
Neuroscience experiment, which uses a python script that extracts brain information and mask from an input image.
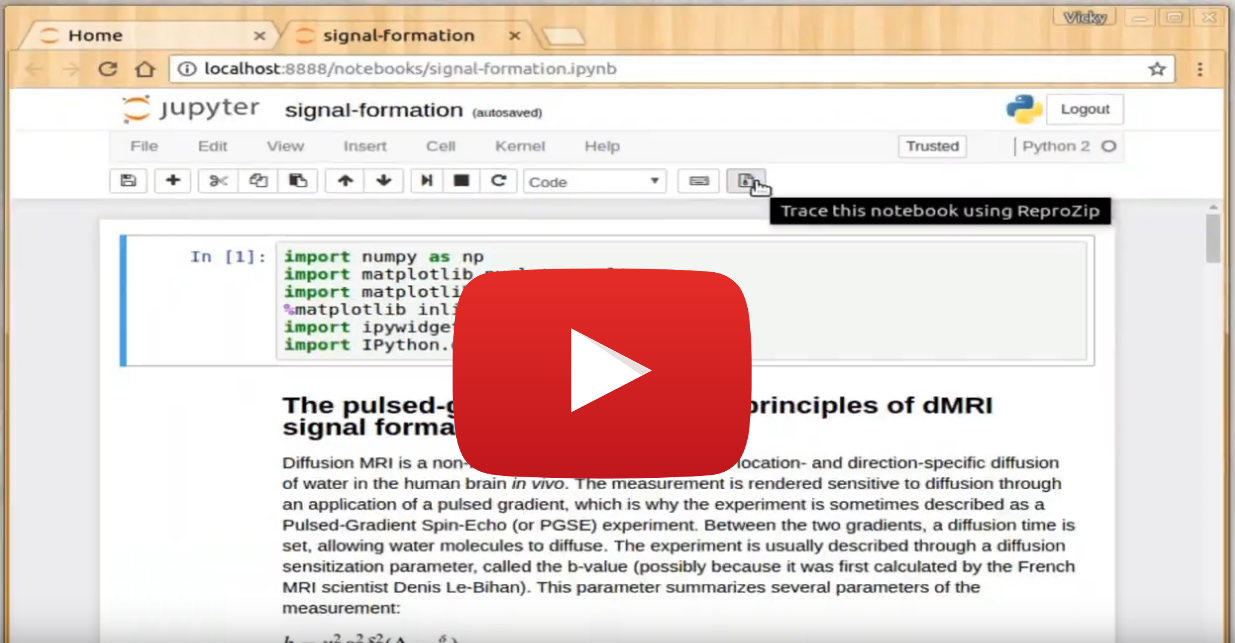
Jupyter Notebook
An example of how to use the reprozip-jupyter plugin to easily capture notebooks with ReproZip.
All the videos showing ReproZip's functionality are on the VIDA-NYU YouTube channel. Various examples of ReproZip packages, including instructions on how to reproduce them, are available in the ReproZip examples website: https://examples.reprozip.org.
Motivation
Reproducibility is a core component of the scientific process: it helps researchers all around the world to verify the results and also to build on them, alowing science to move forward. In natural science, long tradition requires experiments to be described in enough detail so that they can be reproduced by researchers around the world. The same standard, however, has not been widely applied to computational science, where researchers often have to rely on plots, tables, and figures included in papers, which loosely describe the obtained results.
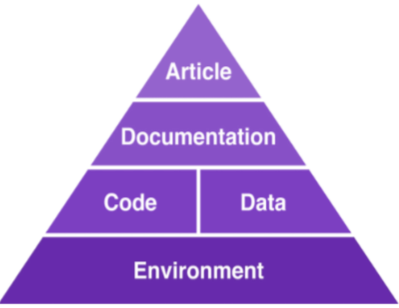
The truth is computational reproducibility can be very painful to achieve for a number of reasons. Take the author-reviewer scenario of a scientific paper as an example. Authors must generate a compendium that encapsulates all the inputs needed to correctly reproduce their experiments: the data, a complete specification of the experiment and its steps, and information about the originating computational environment (OS, hardware architecture, and library dependencies). Keeping track of this information manually is rarely feasible: it is both time-consuming and error-prone. First, computational environments are complex, consisting of many layers of hardware and software, and the configuration of the OS is often hidden. Second, tracking library dependencies is challenging, especially for large experiments. If authors did not plan for it since the beginning of the project, reproducibility is drastically hampered.
For reviewers, even with a compendium in their hands, it may be hard to reproduce the results. There may be no instructions about how to execute the code and explore it further; the experiment may not run on his operating system; there may be missing libraries; library versions may be different; and several issues may arise while trying to install all the required dependencies, a problem colloquially known as dependency hell.
We made ReproZip to try to help alleviate these problems by allowing the user to easily and autoamtically capture all the necessary components in a single, distributable bundle. Also, the tool makes it easier to reproduce an experiment by providing different unpacking methods and interfaces that avoids the need to install all the required dependencies and that makes it possible to run the experiment under different inputs. This provides an easy way to verify and review others' work, bringing reproducibility into the norm of scholarship.
The ReproZip Team
ReproZip is currently being developed at NYU. The team includes:
 Juliana Freire |
 Dennis Shasha |
 Rémi Rampin |
 Fernando Chirigati |
 Vicky Rampin |
 Binal Modi (summer & fall 2018) |
 Khushnaseeb Ali (summer 2018) |
 Zhonheng Li (summer 2017) |
Citing ReproZip
If you wish to cite ReproZip in a paper, please use the following:
ReproZip: Computational Reproducibility With Ease, F. Chirigati, R. Rampin, D. Shasha, and J. Freire. In Proceedings of the 2016 ACM SIGMOD International Conference on Management of Data (SIGMOD), pp. 2085-2088, 2016
The bibtex for this citation can be found here: https://github.com/VIDA-NYU/reprozip/blob/master/CITATION.txt.
A preprint version of this paper is available here. For a complete list of publications, posters, and presentations, please visit the Scholarship page. To reference our website, please use https://www.reprozip.org.
Contact Us
If you'd like to give us feedback, ask questions, or report concerns and issues about ReproZip, you can either open an issue on GitHub or use the mailing-list reprozip@nyu.edu. You can also contribute your use of ReproZip to our reprozip-examples repository.
Contribute to ReproZip!
If you'd like to contribute to the development of ReproZip, please read our CONTRIBUTING.md file in the ReproZip GitHub repository.
We have a number of GitHub issues tagged as help-wanted, question, and good-first-issue. We are hoping that organizing the issues this way will encourage folks of all levels to participate to the development of ReproZip, from those who contribute through discussion around questions, newcomers looking for a good first issue to work on, and developers comfortable looking through a project to write code back.
Acknowledgements
The development of ReproZip has been supported by the Moore and Sloan Foundations, under the NYU Moore-Sloan Data Science Environment.